Phenotypic Mutation 'souris' (pdf version)
| Allele | souris |
|---|
| Mutation Type |
unclassified
(2 bp from exon)
|
|---|
| Chromosome | 13 |
|---|
| Coordinate | 13,857,808 bp (GRCm39) |
|---|
| Base Change | T ⇒ A (forward strand) |
|---|
| Gene |
Lyst
|
| Gene Name | lysosomal trafficking regulator |
|---|
| Synonym(s) | D13Sfk13 |
|---|
| Chromosomal Location |
13,764,982-13,953,388 bp (+) (GRCm39)
|
|---|
| MGI Phenotype |
FUNCTION: [Summary is not available for the mouse gene. This summary is for the human ortholog.] This gene encodes a protein that regulates intracellular protein trafficking in endosomes, and may be involved in pigmentation. Mutations in this gene are associated with Chediak-Higashi syndrome, a lysosomal storage disorder. Alternative splicing results in multiple transcript variants, though the full-length nature of some of these variants has not been determined. [provided by RefSeq, Apr 2013]
PHENOTYPE: Homozygous mice have a phenotype similar to human Chediak-Higashi syndrome patients, exhibiting lysosomal dysfunction with resultant protein storage; diluted coat color; abnormal melanogenesis; immune cell dysfunction resulting in increased susceptibility to bacterial, viral, and parasitic infections and decreased cytotoxic activity against tumor cells. [provided by MGI curators]
|
| Accession Number | NCBI RefSeq: NM_010748; MGI: 107448
|
|---|
| Mapped | Yes |
|---|
| Amino Acid Change |
|
|---|
| Institutional Source | Beutler Lab |
|---|
| Gene Model |
not available |
|---|
| AlphaFold |
no structure available at present |
| SMART Domains |
Protein: ENSMUSP00000106188
Gene: ENSMUSG00000019726
| Domain | Start | End | E-Value | Type |
|---|
|
low complexity region
|
26 |
36 |
N/A |
INTRINSIC |
|
low complexity region
|
72 |
82 |
N/A |
INTRINSIC |
|
low complexity region
|
399 |
412 |
N/A |
INTRINSIC |
|
low complexity region
|
1333 |
1344 |
N/A |
INTRINSIC |
|
low complexity region
|
2295 |
2307 |
N/A |
INTRINSIC |
|
low complexity region
|
2427 |
2445 |
N/A |
INTRINSIC |
|
low complexity region
|
2534 |
2546 |
N/A |
INTRINSIC |
|
Pfam:PH_BEACH
|
3006 |
3101 |
5.8e-25 |
PFAM |
|
Beach
|
3118 |
3408 |
1.25e-193 |
SMART |
|
Blast:Beach
|
3441 |
3478 |
9e-13 |
BLAST |
|
WD40
|
3539 |
3579 |
5.75e-1 |
SMART |
|
WD40
|
3591 |
3630 |
2.89e-5 |
SMART |
|
WD40
|
3633 |
3676 |
1.38e0 |
SMART |
|
WD40
|
3724 |
3765 |
1.27e-1 |
SMART |
|
|---|
| Predicted Effect |
probably benign
|
|---|
| Predicted Effect |
probably benign
|
|---|
| Meta Mutation Damage Score |
Not available  |
|---|
| Is this an essential gene? |
Possibly nonessential (E-score: 0.395)  |
|---|
| Phenotypic Category |
Autosomal Recessive |
| Candidate Explorer Status |
loading ... |
Single pedigree
Linkage Analysis Data
|
|
| Penetrance | 100% |
|---|
| Alleles Listed at MGI | All alleles(45) : Gene trapped(30) Spontaneous(8) Chemically induced(6) Radiation induced(1)
|
|---|
| Lab Alleles |
| Allele | Source | Chr | Coord | Type | Predicted Effect | PPH Score |
|---|
|
IGL00156:Lyst
|
APN |
13 |
13823463 |
missense |
probably benign |
|
|
IGL00474:Lyst
|
APN |
13 |
13818121 |
missense |
possibly damaging |
0.48 |
|
IGL00484:Lyst
|
APN |
13 |
13884188 |
missense |
probably benign |
0.02 |
|
IGL00492:Lyst
|
APN |
13 |
13852760 |
missense |
possibly damaging |
0.54 |
|
IGL00807:Lyst
|
APN |
13 |
13825008 |
missense |
possibly damaging |
0.91 |
|
IGL00949:Lyst
|
APN |
13 |
13810070 |
missense |
possibly damaging |
0.87 |
|
IGL00952:Lyst
|
APN |
13 |
13852692 |
missense |
probably benign |
0.05 |
|
IGL01305:Lyst
|
APN |
13 |
13852641 |
missense |
probably benign |
0.01 |
|
IGL01317:Lyst
|
APN |
13 |
13845455 |
missense |
probably benign |
|
|
IGL01419:Lyst
|
APN |
13 |
13810423 |
missense |
probably benign |
0.00 |
|
IGL01445:Lyst
|
APN |
13 |
13826299 |
missense |
probably benign |
0.00 |
|
IGL01690:Lyst
|
APN |
13 |
13917831 |
missense |
probably damaging |
1.00 |
|
IGL01791:Lyst
|
APN |
13 |
13809887 |
missense |
probably damaging |
1.00 |
|
IGL01809:Lyst
|
APN |
13 |
13812388 |
missense |
probably damaging |
1.00 |
|
IGL01896:Lyst
|
APN |
13 |
13810162 |
missense |
probably benign |
0.04 |
|
IGL01938:Lyst
|
APN |
13 |
13812009 |
missense |
possibly damaging |
0.93 |
|
IGL01986:Lyst
|
APN |
13 |
13950212 |
critical splice donor site |
probably null |
|
|
IGL02022:Lyst
|
APN |
13 |
13838629 |
nonsense |
probably null |
|
|
IGL02044:Lyst
|
APN |
13 |
13887431 |
missense |
probably damaging |
1.00 |
|
IGL02157:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02185:Lyst
|
APN |
13 |
13835678 |
nonsense |
probably null |
|
|
IGL02215:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02245:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02246:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02247:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02297:Lyst
|
APN |
13 |
13812677 |
nonsense |
probably null |
|
|
IGL02411:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02415:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02419:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02420:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02429:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02501:Lyst
|
APN |
13 |
13886230 |
missense |
probably benign |
0.02 |
|
IGL02522:Lyst
|
APN |
13 |
13809290 |
missense |
possibly damaging |
0.81 |
|
IGL02535:Lyst
|
APN |
13 |
13824927 |
missense |
probably benign |
0.00 |
|
IGL02596:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02601:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02603:Lyst
|
APN |
13 |
13835541 |
missense |
probably benign |
|
|
IGL02608:Lyst
|
APN |
13 |
13887339 |
missense |
probably damaging |
0.98 |
|
IGL02622:Lyst
|
APN |
13 |
13855975 |
missense |
probably damaging |
1.00 |
|
IGL02690:Lyst
|
APN |
13 |
13815710 |
missense |
possibly damaging |
0.58 |
|
IGL02715:Lyst
|
APN |
13 |
13848905 |
splice site |
probably null |
|
|
IGL02725:Lyst
|
APN |
13 |
13935412 |
missense |
probably damaging |
1.00 |
|
IGL02729:Lyst
|
APN |
13 |
13921194 |
missense |
possibly damaging |
0.95 |
|
IGL02729:Lyst
|
APN |
13 |
13848924 |
missense |
possibly damaging |
0.81 |
|
IGL02820:Lyst
|
APN |
13 |
13812643 |
missense |
probably benign |
0.03 |
|
IGL02945:Lyst
|
APN |
13 |
13935783 |
missense |
possibly damaging |
0.48 |
|
IGL02981:Lyst
|
APN |
13 |
13809496 |
missense |
probably damaging |
0.99 |
|
IGL03087:Lyst
|
APN |
13 |
13809641 |
missense |
probably damaging |
1.00 |
|
IGL03149:Lyst
|
APN |
13 |
13856029 |
missense |
probably benign |
0.14 |
|
IGL03158:Lyst
|
APN |
13 |
13826337 |
critical splice donor site |
probably null |
|
|
IGL03226:Lyst
|
APN |
13 |
13884144 |
missense |
probably benign |
0.01 |
|
IGL03242:Lyst
|
APN |
13 |
13831466 |
nonsense |
probably null |
|
|
IGL03385:Lyst
|
APN |
13 |
13831565 |
nonsense |
probably null |
|
|
50-cal
|
UTSW |
13 |
13882797 |
critical splice donor site |
probably null |
|
|
charcoal
|
UTSW |
13 |
13871346 |
nonsense |
probably null |
|
|
charlotte_gray
|
UTSW |
13 |
13602026 |
intron |
probably benign |
|
|
charzard
|
UTSW |
13 |
13821668 |
nonsense |
probably null |
|
|
grey_wolf
|
UTSW |
13 |
|
unclassified |
|
|
|
lightspeed
|
UTSW |
13 |
13915121 |
missense |
possibly damaging |
0.91 |
|
pardon
|
UTSW |
13 |
13852537 |
missense |
probably benign |
0.00 |
|
robin
|
UTSW |
13 |
13823387 |
nonsense |
probably null |
|
|
sooty
|
UTSW |
13 |
|
unclassified |
|
|
|
Swallow
|
UTSW |
13 |
13932007 |
missense |
probably benign |
0.00 |
|
vulpix
|
UTSW |
13 |
13871379 |
splice site |
probably null |
|
|
ANU22:Lyst
|
UTSW |
13 |
13852641 |
missense |
probably benign |
0.01 |
|
IGL02835:Lyst
|
UTSW |
13 |
13835685 |
missense |
possibly damaging |
0.82 |
|
P0031:Lyst
|
UTSW |
13 |
13838616 |
missense |
probably damaging |
1.00 |
|
R0012:Lyst
|
UTSW |
13 |
13862279 |
missense |
probably benign |
0.10 |
|
R0012:Lyst
|
UTSW |
13 |
13862279 |
missense |
probably benign |
0.10 |
|
R0031:Lyst
|
UTSW |
13 |
13882741 |
missense |
probably benign |
0.14 |
|
R0115:Lyst
|
UTSW |
13 |
13852537 |
missense |
probably benign |
0.00 |
|
R0212:Lyst
|
UTSW |
13 |
13810570 |
missense |
possibly damaging |
0.93 |
|
R0386:Lyst
|
UTSW |
13 |
13882799 |
splice site |
probably benign |
|
|
R0393:Lyst
|
UTSW |
13 |
13821664 |
missense |
probably benign |
0.01 |
|
R0415:Lyst
|
UTSW |
13 |
13886195 |
splice site |
probably benign |
|
|
R0446:Lyst
|
UTSW |
13 |
13812633 |
missense |
probably benign |
0.00 |
|
R0481:Lyst
|
UTSW |
13 |
13852537 |
missense |
probably benign |
0.00 |
|
R0499:Lyst
|
UTSW |
13 |
13791298 |
missense |
probably damaging |
1.00 |
|
R0506:Lyst
|
UTSW |
13 |
13812600 |
missense |
probably benign |
|
|
R0530:Lyst
|
UTSW |
13 |
13931891 |
splice site |
probably benign |
|
|
R0541:Lyst
|
UTSW |
13 |
13855878 |
missense |
probably benign |
0.00 |
|
R0570:Lyst
|
UTSW |
13 |
13883971 |
missense |
probably benign |
0.26 |
|
R0680:Lyst
|
UTSW |
13 |
13824926 |
missense |
probably benign |
0.01 |
|
R0842:Lyst
|
UTSW |
13 |
13852826 |
nonsense |
probably null |
|
|
R0848:Lyst
|
UTSW |
13 |
13809515 |
missense |
probably benign |
0.00 |
|
R1014:Lyst
|
UTSW |
13 |
13808645 |
missense |
possibly damaging |
0.49 |
|
R1205:Lyst
|
UTSW |
13 |
13854787 |
missense |
probably benign |
|
|
R1251:Lyst
|
UTSW |
13 |
13809068 |
missense |
probably benign |
0.00 |
|
R1304:Lyst
|
UTSW |
13 |
13926569 |
nonsense |
probably null |
|
|
R1398:Lyst
|
UTSW |
13 |
13915121 |
missense |
possibly damaging |
0.91 |
|
R1445:Lyst
|
UTSW |
13 |
13814639 |
missense |
possibly damaging |
0.94 |
|
R1475:Lyst
|
UTSW |
13 |
13882797 |
critical splice donor site |
probably null |
|
|
R1479:Lyst
|
UTSW |
13 |
13809067 |
missense |
probably benign |
0.00 |
|
R1484:Lyst
|
UTSW |
13 |
13852775 |
missense |
probably benign |
0.01 |
|
R1498:Lyst
|
UTSW |
13 |
13824960 |
missense |
possibly damaging |
0.49 |
|
R1540:Lyst
|
UTSW |
13 |
13809686 |
missense |
possibly damaging |
0.81 |
|
R1611:Lyst
|
UTSW |
13 |
13809482 |
missense |
probably damaging |
0.97 |
|
R1653:Lyst
|
UTSW |
13 |
13809811 |
missense |
probably damaging |
1.00 |
|
R1669:Lyst
|
UTSW |
13 |
13818672 |
missense |
possibly damaging |
0.90 |
|
R1686:Lyst
|
UTSW |
13 |
13809290 |
missense |
possibly damaging |
0.81 |
|
R1694:Lyst
|
UTSW |
13 |
13835746 |
missense |
probably damaging |
0.98 |
|
R1747:Lyst
|
UTSW |
13 |
13932007 |
missense |
probably benign |
0.00 |
|
R1793:Lyst
|
UTSW |
13 |
13821668 |
nonsense |
probably null |
|
|
R1871:Lyst
|
UTSW |
13 |
13826297 |
missense |
probably benign |
0.00 |
|
R1905:Lyst
|
UTSW |
13 |
13808719 |
missense |
probably benign |
|
|
R1958:Lyst
|
UTSW |
13 |
13791203 |
missense |
probably damaging |
1.00 |
|
R1969:Lyst
|
UTSW |
13 |
13904929 |
missense |
probably damaging |
0.99 |
|
R2040:Lyst
|
UTSW |
13 |
13815807 |
missense |
probably benign |
0.00 |
|
R2109:Lyst
|
UTSW |
13 |
13887405 |
missense |
possibly damaging |
0.46 |
|
R2116:Lyst
|
UTSW |
13 |
13810286 |
missense |
probably damaging |
0.99 |
|
R2121:Lyst
|
UTSW |
13 |
13835556 |
missense |
probably damaging |
1.00 |
|
R2127:Lyst
|
UTSW |
13 |
13809847 |
missense |
probably damaging |
1.00 |
|
R2187:Lyst
|
UTSW |
13 |
13883926 |
missense |
possibly damaging |
0.61 |
|
R2238:Lyst
|
UTSW |
13 |
13917848 |
missense |
probably benign |
0.41 |
|
R2258:Lyst
|
UTSW |
13 |
13812243 |
missense |
probably benign |
0.00 |
|
R2292:Lyst
|
UTSW |
13 |
13915080 |
missense |
probably damaging |
1.00 |
|
R2368:Lyst
|
UTSW |
13 |
13871248 |
missense |
probably damaging |
0.96 |
|
R2908:Lyst
|
UTSW |
13 |
13844458 |
missense |
probably benign |
0.03 |
|
R3001:Lyst
|
UTSW |
13 |
13871290 |
missense |
probably benign |
|
|
R3002:Lyst
|
UTSW |
13 |
13871290 |
missense |
probably benign |
|
|
R3024:Lyst
|
UTSW |
13 |
13833272 |
missense |
probably benign |
|
|
R3113:Lyst
|
UTSW |
13 |
13844512 |
missense |
probably benign |
0.12 |
|
R3406:Lyst
|
UTSW |
13 |
13809815 |
missense |
possibly damaging |
0.56 |
|
R3972:Lyst
|
UTSW |
13 |
13881210 |
missense |
possibly damaging |
0.67 |
|
R3978:Lyst
|
UTSW |
13 |
13808753 |
missense |
possibly damaging |
0.82 |
|
R4032:Lyst
|
UTSW |
13 |
13791250 |
missense |
probably damaging |
1.00 |
|
R4192:Lyst
|
UTSW |
13 |
13915098 |
missense |
probably damaging |
1.00 |
|
R4206:Lyst
|
UTSW |
13 |
13810574 |
missense |
probably benign |
0.03 |
|
R4298:Lyst
|
UTSW |
13 |
13809472 |
missense |
probably damaging |
1.00 |
|
R4344:Lyst
|
UTSW |
13 |
13873051 |
missense |
probably benign |
0.06 |
|
R4441:Lyst
|
UTSW |
13 |
13809968 |
missense |
probably damaging |
1.00 |
|
R4445:Lyst
|
UTSW |
13 |
13884149 |
missense |
probably benign |
0.42 |
|
R4477:Lyst
|
UTSW |
13 |
13809968 |
missense |
probably damaging |
1.00 |
|
R4493:Lyst
|
UTSW |
13 |
13809968 |
missense |
probably damaging |
1.00 |
|
R4494:Lyst
|
UTSW |
13 |
13809968 |
missense |
probably damaging |
1.00 |
|
R4495:Lyst
|
UTSW |
13 |
13809968 |
missense |
probably damaging |
1.00 |
|
R4622:Lyst
|
UTSW |
13 |
13848983 |
missense |
probably benign |
0.01 |
|
R4638:Lyst
|
UTSW |
13 |
13871379 |
splice site |
probably null |
|
|
R4658:Lyst
|
UTSW |
13 |
13809968 |
missense |
probably damaging |
1.00 |
|
R4675:Lyst
|
UTSW |
13 |
13809968 |
missense |
probably damaging |
1.00 |
|
R4719:Lyst
|
UTSW |
13 |
13824935 |
missense |
probably benign |
|
|
R4729:Lyst
|
UTSW |
13 |
13812486 |
missense |
probably damaging |
1.00 |
|
R4774:Lyst
|
UTSW |
13 |
13915182 |
missense |
probably damaging |
1.00 |
|
R4811:Lyst
|
UTSW |
13 |
13951685 |
missense |
probably benign |
0.33 |
|
R4877:Lyst
|
UTSW |
13 |
13857734 |
missense |
probably damaging |
1.00 |
|
R4920:Lyst
|
UTSW |
13 |
13821645 |
missense |
possibly damaging |
0.79 |
|
R4933:Lyst
|
UTSW |
13 |
13933963 |
missense |
probably benign |
0.12 |
|
R4933:Lyst
|
UTSW |
13 |
13812349 |
missense |
probably damaging |
0.98 |
|
R4958:Lyst
|
UTSW |
13 |
13810048 |
missense |
probably benign |
0.00 |
|
R4982:Lyst
|
UTSW |
13 |
13900539 |
missense |
probably damaging |
1.00 |
|
R4992:Lyst
|
UTSW |
13 |
13835748 |
missense |
probably damaging |
1.00 |
|
R5024:Lyst
|
UTSW |
13 |
13808989 |
missense |
probably benign |
|
|
R5049:Lyst
|
UTSW |
13 |
13810649 |
missense |
probably damaging |
1.00 |
|
R5079:Lyst
|
UTSW |
13 |
13931938 |
missense |
probably benign |
0.08 |
|
R5254:Lyst
|
UTSW |
13 |
13857655 |
missense |
probably benign |
0.00 |
|
R5266:Lyst
|
UTSW |
13 |
13835555 |
missense |
probably damaging |
1.00 |
|
R5279:Lyst
|
UTSW |
13 |
13823387 |
nonsense |
probably null |
|
|
R5285:Lyst
|
UTSW |
13 |
13809011 |
missense |
probably benign |
0.01 |
|
R5364:Lyst
|
UTSW |
13 |
13831439 |
missense |
probably benign |
0.35 |
|
R5435:Lyst
|
UTSW |
13 |
13951649 |
missense |
possibly damaging |
0.64 |
|
R5516:Lyst
|
UTSW |
13 |
13818707 |
missense |
probably benign |
0.10 |
|
R5524:Lyst
|
UTSW |
13 |
13921364 |
missense |
probably benign |
0.03 |
|
R5591:Lyst
|
UTSW |
13 |
13917918 |
missense |
probably damaging |
0.99 |
|
R5592:Lyst
|
UTSW |
13 |
13917918 |
missense |
probably damaging |
0.99 |
|
R5593:Lyst
|
UTSW |
13 |
13917918 |
missense |
probably damaging |
0.99 |
|
R5594:Lyst
|
UTSW |
13 |
13917918 |
missense |
probably damaging |
0.99 |
|
R5594:Lyst
|
UTSW |
13 |
13933982 |
missense |
probably benign |
0.00 |
|
R5644:Lyst
|
UTSW |
13 |
13812081 |
missense |
possibly damaging |
0.58 |
|
R5659:Lyst
|
UTSW |
13 |
13809212 |
missense |
possibly damaging |
0.58 |
|
R5741:Lyst
|
UTSW |
13 |
13808615 |
missense |
probably benign |
0.44 |
|
R5908:Lyst
|
UTSW |
13 |
13871346 |
nonsense |
probably null |
|
|
R5969:Lyst
|
UTSW |
13 |
13862398 |
splice site |
probably null |
|
|
R6128:Lyst
|
UTSW |
13 |
13933964 |
missense |
possibly damaging |
0.67 |
|
R6271:Lyst
|
UTSW |
13 |
13833339 |
missense |
probably benign |
0.30 |
|
R6315:Lyst
|
UTSW |
13 |
13818089 |
missense |
probably benign |
|
|
R6318:Lyst
|
UTSW |
13 |
13917896 |
missense |
possibly damaging |
0.88 |
|
R6555:Lyst
|
UTSW |
13 |
13823510 |
missense |
probably benign |
0.01 |
|
R6663:Lyst
|
UTSW |
13 |
13838701 |
splice site |
probably null |
|
|
R6701:Lyst
|
UTSW |
13 |
13856070 |
missense |
probably benign |
0.06 |
|
R6711:Lyst
|
UTSW |
13 |
13809820 |
missense |
possibly damaging |
0.80 |
|
R6909:Lyst
|
UTSW |
13 |
13917960 |
missense |
probably damaging |
1.00 |
|
R6915:Lyst
|
UTSW |
13 |
13900629 |
missense |
probably benign |
0.01 |
|
R6929:Lyst
|
UTSW |
13 |
13917909 |
missense |
probably damaging |
1.00 |
|
R6960:Lyst
|
UTSW |
13 |
13808663 |
missense |
probably benign |
0.12 |
|
R7018:Lyst
|
UTSW |
13 |
13918044 |
critical splice donor site |
probably null |
|
|
R7037:Lyst
|
UTSW |
13 |
13791251 |
missense |
probably damaging |
1.00 |
|
R7045:Lyst
|
UTSW |
13 |
13812293 |
missense |
probably damaging |
1.00 |
|
R7045:Lyst
|
UTSW |
13 |
13809485 |
missense |
probably benign |
0.34 |
|
R7070:Lyst
|
UTSW |
13 |
13932029 |
missense |
probably benign |
0.23 |
|
R7188:Lyst
|
UTSW |
13 |
13926675 |
missense |
possibly damaging |
0.66 |
|
R7201:Lyst
|
UTSW |
13 |
13883885 |
nonsense |
probably null |
|
|
R7210:Lyst
|
UTSW |
13 |
13831568 |
missense |
probably damaging |
1.00 |
|
R7229:Lyst
|
UTSW |
13 |
13818094 |
missense |
probably benign |
0.00 |
|
R7293:Lyst
|
UTSW |
13 |
13854822 |
missense |
probably benign |
0.01 |
|
R7318:Lyst
|
UTSW |
13 |
13932028 |
missense |
probably benign |
0.13 |
|
R7344:Lyst
|
UTSW |
13 |
13881140 |
missense |
probably benign |
|
|
R7426:Lyst
|
UTSW |
13 |
13812109 |
missense |
probably benign |
|
|
R7522:Lyst
|
UTSW |
13 |
13821668 |
nonsense |
probably null |
|
|
R7583:Lyst
|
UTSW |
13 |
13810472 |
missense |
probably damaging |
1.00 |
|
R7606:Lyst
|
UTSW |
13 |
13812060 |
missense |
probably damaging |
1.00 |
|
R7636:Lyst
|
UTSW |
13 |
13791332 |
critical splice donor site |
probably null |
|
|
R7658:Lyst
|
UTSW |
13 |
13905061 |
missense |
possibly damaging |
0.63 |
|
R7685:Lyst
|
UTSW |
13 |
13844450 |
missense |
probably benign |
0.00 |
|
R7689:Lyst
|
UTSW |
13 |
13857808 |
critical splice donor site |
probably null |
|
|
R7765:Lyst
|
UTSW |
13 |
13884117 |
missense |
possibly damaging |
0.75 |
|
R7779:Lyst
|
UTSW |
13 |
13809128 |
missense |
probably damaging |
1.00 |
|
R7871:Lyst
|
UTSW |
13 |
13810637 |
nonsense |
probably null |
|
|
R7872:Lyst
|
UTSW |
13 |
13810450 |
missense |
probably benign |
0.14 |
|
R7884:Lyst
|
UTSW |
13 |
13882268 |
missense |
probably benign |
0.09 |
|
R7890:Lyst
|
UTSW |
13 |
13915154 |
missense |
probably damaging |
0.99 |
|
R7916:Lyst
|
UTSW |
13 |
13821657 |
missense |
possibly damaging |
0.64 |
|
R7948:Lyst
|
UTSW |
13 |
13921174 |
missense |
possibly damaging |
0.59 |
|
R7956:Lyst
|
UTSW |
13 |
13815788 |
missense |
possibly damaging |
0.80 |
|
R8048:Lyst
|
UTSW |
13 |
13862230 |
missense |
probably benign |
0.12 |
|
R8085:Lyst
|
UTSW |
13 |
13808894 |
missense |
probably damaging |
0.98 |
|
R8165:Lyst
|
UTSW |
13 |
13872945 |
missense |
probably damaging |
0.99 |
|
R8235:Lyst
|
UTSW |
13 |
13935323 |
missense |
possibly damaging |
0.69 |
|
R8237:Lyst
|
UTSW |
13 |
13826317 |
missense |
probably benign |
0.00 |
|
R8275:Lyst
|
UTSW |
13 |
13950667 |
missense |
probably benign |
0.02 |
|
R8300:Lyst
|
UTSW |
13 |
13838643 |
missense |
possibly damaging |
0.79 |
|
R8350:Lyst
|
UTSW |
13 |
13824973 |
nonsense |
probably null |
|
|
R8526:Lyst
|
UTSW |
13 |
13935391 |
missense |
probably damaging |
0.99 |
|
R8551:Lyst
|
UTSW |
13 |
13808645 |
missense |
possibly damaging |
0.77 |
|
R8723:Lyst
|
UTSW |
13 |
13887342 |
missense |
possibly damaging |
0.89 |
|
R8772:Lyst
|
UTSW |
13 |
13812077 |
nonsense |
probably null |
|
|
R8778:Lyst
|
UTSW |
13 |
13903152 |
missense |
possibly damaging |
0.89 |
|
R8778:Lyst
|
UTSW |
13 |
13810361 |
missense |
possibly damaging |
0.89 |
|
R8801:Lyst
|
UTSW |
13 |
13835595 |
missense |
probably benign |
0.10 |
|
R8837:Lyst
|
UTSW |
13 |
13852548 |
missense |
probably benign |
|
|
R8874:Lyst
|
UTSW |
13 |
13812147 |
missense |
probably benign |
|
|
R8878:Lyst
|
UTSW |
13 |
13815661 |
missense |
probably benign |
0.00 |
|
R8891:Lyst
|
UTSW |
13 |
13887435 |
missense |
possibly damaging |
0.67 |
|
R9077:Lyst
|
UTSW |
13 |
13857693 |
missense |
probably benign |
0.02 |
|
R9127:Lyst
|
UTSW |
13 |
13808827 |
missense |
probably damaging |
1.00 |
|
R9143:Lyst
|
UTSW |
13 |
13835750 |
missense |
probably damaging |
0.98 |
|
R9216:Lyst
|
UTSW |
13 |
13823188 |
missense |
probably benign |
|
|
R9217:Lyst
|
UTSW |
13 |
13871245 |
missense |
probably benign |
0.01 |
|
R9291:Lyst
|
UTSW |
13 |
13883938 |
missense |
probably benign |
0.01 |
|
R9302:Lyst
|
UTSW |
13 |
13904947 |
missense |
possibly damaging |
0.46 |
|
R9370:Lyst
|
UTSW |
13 |
13935333 |
missense |
probably damaging |
1.00 |
|
R9402:Lyst
|
UTSW |
13 |
13812463 |
missense |
probably benign |
|
|
R9457:Lyst
|
UTSW |
13 |
13862330 |
missense |
possibly damaging |
0.83 |
|
R9481:Lyst
|
UTSW |
13 |
13857653 |
missense |
possibly damaging |
0.68 |
|
R9563:Lyst
|
UTSW |
13 |
13812408 |
missense |
probably benign |
0.36 |
|
R9623:Lyst
|
UTSW |
13 |
13852587 |
missense |
probably benign |
|
|
R9661:Lyst
|
UTSW |
13 |
13808779 |
missense |
probably benign |
0.01 |
|
R9682:Lyst
|
UTSW |
13 |
13831526 |
missense |
probably benign |
0.21 |
|
R9743:Lyst
|
UTSW |
13 |
13809323 |
missense |
possibly damaging |
0.67 |
|
R9801:Lyst
|
UTSW |
13 |
13809290 |
missense |
probably damaging |
0.97 |
|
RF001:Lyst
|
UTSW |
13 |
13810426 |
missense |
probably benign |
|
|
RF002:Lyst
|
UTSW |
13 |
13808948 |
missense |
probably benign |
0.05 |
|
X0024:Lyst
|
UTSW |
13 |
13809033 |
missense |
probably benign |
0.00 |
|
X0026:Lyst
|
UTSW |
13 |
13926555 |
missense |
probably damaging |
0.99 |
|
Z1088:Lyst
|
UTSW |
13 |
13918018 |
missense |
probably benign |
0.09 |
|
Z1176:Lyst
|
UTSW |
13 |
13951664 |
missense |
probably benign |
0.27 |
|
Z1176:Lyst
|
UTSW |
13 |
13814692 |
missense |
probably damaging |
1.00 |
|
Z1177:Lyst
|
UTSW |
13 |
13854719 |
missense |
possibly damaging |
0.73 |
|
|---|
| Mode of Inheritance |
Autosomal Recessive |
| Local Stock | Embryos, Sperm, gDNA |
|---|
| MMRRC Submission |
010470-UCD
|
|---|
| Last Updated |
2018-08-02 11:40 AM
by Diantha La Vine
|
| Record Created |
unknown
|
| Record Posted |
2007-10-10 |
|---|
| Phenotypic Description |
The souris phenotype was identified in two ENU-induced G3 littermates. Souris mice have a uniform dark gray coat color. The ears, feet and tail have a light pink to white color. The souris mutation confers mouse cytomegalovirus (MCMV) susceptibility, with homozygotes exhibiting a high splenic viral titer 5 days post-infection (similar to MCMV-infected Balb/c mice) ( MCMV Susceptibility and Resistance Screen). Souris mice are susceptible to infection with Listeria monocytogenes. NK cells from souris mice fail to degranulate after antibody stimulation of NKp46 or Ly49H receptors, or after exposure to YAC-1 cells or PMA/ionomycin. However, intracellular production of interferon (IFN)-γ is normal after PMA/ionomycin stimulation.
Complementation testing demonstrated that the souris mutation is allelic with the beige locus, in which a Lyst mutation was identified ( 1;2). |
|---|
| Nature of Mutation |  The souris mutation was mapped to Chromosome 13, and corresponds to a T to A transversion in the donor splice site of intron 27 (G TAAG -> G AAAG) of the Lyst gene (position 92815 in the Genbank genomic region NC_000079 for linear genomic DNA sequence of Lyst). As Lyst cDNA from souris mice has not been sequenced, it is unknown how the mutation affects processing of the Lyst transcript. The mutation likely results in skipping of the 167-nucleotide exon 27 (out of 53 total exons), destroying the reading frame (aberrant amino acids after position 2474), and creating a premature stop codon that would truncate the protein after amino acid 2482:
<-- exon 26 <--exon 27 intron 27--> exon 28-->
91104 TTAGAAAAGAA……AGGACACAAA GTAAGAATG……………ATATGGCTTTGGCCCTGCAGCTTAG 93574
2472 -L--E--K--N……-R--T--Q-- --Y--G--F--F--P--A--A--* 2482
correct deleted aberrant
The donor splice site of intron 27, which is destroyed by the souris mutation, is shown in blue; the mutated nucleotide is shown in red.
|
|---|
Illustration of Mutations in
Gene & Protein |
|
|---|
| Protein Prediction |
The Lyst gene encodes the protein Lyst (also CHS/Beige), a 3788-amino acid protein whose biochemical functions remain unknown (Figure 1). A large N-terminal portion of the protein (amino acids 1-3132) contains approximately twenty repeats with homology to ARM (Armadillo) and HEAT ( huntingtin, elongation factor 3, A subunit of protein phosphatase A, target of rapamycin) repeat motifs ( 3;4). ARM and HEAT motifs are α-helical domains of about 50 amino acids that pack together to form elongated “solenoids” ( 5); evidence suggests they mediate protein associations at the membrane ( 6), and vesicle transport ( 7), respectively. A perilipin domain (amino acids 1079-1313) may interact with lipids, as has been suggested for several other proteins ( 8). A 674-amino acid construct containing the perilipin domain can function as a dominant negative peptide that causes enlarged lysosomes when overexpressed in Cos-7 or HeLa cells ( 9). The C-terminus of Lyst contains two distinct domains, a BEACH ( beige and chediak) domain (amino acids 3132-3472) and seven WD40 motifs ( 3). The BEACH domain is a 345-amino acid region of unknown function ( 3), and WD40 motifs are protein interaction motifs that typically form β sheets arranged in a 7-bladed β propeller fold ( 10). Database searches have identified one or more proteins in each of S. cerevisiae, C. elegans, D. discoideum, D. melanogaster, mice and humans that contain both BEACH and WD40 domains ( 3;4). So far, these proteins appear to have diverse functions [reviewed in ( 4)]. Notably, a C-terminal fragment of Lyst containing the BEACH and WD40 domains can act dominant negatively to cause enlarged lysosomes when expressed in Cos-7 or HeLa cells ( 9).
In addition to these protein domains, Lyst also contains potential phosphorylation sites for protein kinase C (PKC), casein kinase II (CKII), c-AMP-dependent protein kinase, and a tyrosine kinase ( 1).
The souris mutation results in eight aberrant amino acids after position 2474, followed by premature truncation of the protein. This would delete four of the ARM/HEAT motifs, the BEACH domain and the WD40 motifs. Expression of the mutated Lyst protein has not been tested.
|
|---|
| Expression/Localization | Lyst transcripts are expressed ubiquitously in both mouse and human tissues ( 1;2). Several alternative transcripts are detected in various tissues, but their functional differences are unknown ( 1;11). The Lyst protein is localized in the cytoplasm ( 9;12).
|
|---|
| Background |
In humans, mutations in the Lyst gene cause Chediak-Higashi Syndrome (CHS, OMIM #214500), a rare autosomal recessive disorder characterized by oculocutaneous albinism, severe immune deficiency, bleeding tendency, recurrent pyogenic infection, progressive neurologic defects and a lymphoproliferative syndrome [( 1;2), reviewed in ( 4)]. These defects are caused by the aberrant formation of giant granules within a variety of cell types, and disrupted intracellular protein trafficking ( 4;13;14). The enlarged granules consist of organelles such as lysosomes, melanosomes, cytolytic granules and platelet dense bodies, and it is thought that the increased size of these organelles inhibits their migration and fusion at the cell surface and/or organelle-organelle fusion (Figure 2). For example, CHS melanocytes produce normal amounts of melanin, the pigment for skin, hair and eyes, but melanosomes are abnormally large and fail to release their contents for absorption by keratinocytes ( 15). Secretion of abnormally large platelet dense bodies is delayed in CHS platelets, resulting in defective platelet coagulation and the bleeding tendency of CHS patients ( 16;17).
Defective lysosome-related functions in immune cells lead to immune deficiency, recurrent bacterial infections and lymphoproliferative disorder in CHS patients. CHS macrophages and polymorphonuclear leukocytes have normal phagocytic ability, but delayed fusion of phagosomes with lysosomes, allowing bacterial replication and escape and leading to persistent infections ( 4). Cytolytic T cells (CTLs) from CHS patients also fail to kill target cells recognized by the T cell receptor (TCR) due to an inability to secrete granules containing lytic proteins ( 18). Although these granules are abnormally large in CHS CTLs, they contain normal levels of properly processed lytic proteins ( 18;19). CHS patients also have impaired NK cell function, presumably due to defective degranulation of the single large granule, rather than the normal multiple small granules, present in these cells ( 20;21). Patients may also display an ‘accelerated phase’ of the disease, in which activated T cells infiltrate the major organs of the body; this phase may be associated with viral infection ( 22).
Human patients with CHS usually die during childhood from the immunologic complications of the disease. Bone marrow transplantation is an effective treatment to extend the lifespan of patients ( 23;24), although it does not prevent the extrahematopoietic symptoms of CHS. Humans with CHS that receive bone marrow transplant and live into adulthood develop neurologic defects including ataxia, sensory deficits and neurodegeneration ( 25). These are thought to result from accumulation of cytoplasmic inclusions in neurons that may prevent normal synaptic transmission ( 26).
In mice, mutations in Lyst cause the beige phenotype ( 1;2). As in humans, beige mice exhibit hypopigmentation, bleeding tendency, and defective immune cell function resulting from the formation of giant granules in melanosomes, lymphocytes, neutrophils, and other cell types ( 14;27;28). Beige mice have defective NK cell ( 29) and CTL function ( 30), and increased susceptibility to infections ( 31;32). However, beige mice do not develop lymphoproliferative disorder, even after challenge with infection ( 31). |
|---|
| Putative Mechanism | There is no clear understanding of the molecular mechanisms of Lyst protein function, or how its loss leads to the formation of enlarged lysosomes and lysosome-related organelles. Overexpression of Lyst in cultured mouse fibroblasts results in reduced lysosome size, suggesting that Lyst is important for vesicle fission ( 12). On the other hand, a yeast-two hybrid screen of cDNA libraries from human heart, keratinocyte, fetal brain, fetal liver and fetal kidney identified Lyst interactions with several regulators of the SNARE [ SNAP (soluble NSF attachment protein) receptor] complex, which mediates vesicle fusion with the cell membrane or with target compartments such as lysosomes ( 33). Although the interactions were tested in vitro and have yet to be confirmed genetically, they raise the possibility that Lyst regulates SNARE-mediated membrane fusion. In support of this hypothesis, lysosomal exocytosis and membrane resealing are impaired after membrane wounding in CHS or beige fibroblasts ( 34). Lysosomal exocytosis contributes to the repair of plasma membrane lesions in a Ca 2+-dependent manner. Furthermore, survival of membrane-wounded CHS and beige fibroblasts is reduced compared to wild type, and correlates with levels of lysosomal exocytosis ( 34). Thus, these data suggest that Lyst promotes both the processes of vesicle budding and vesicle fusion.
Unfortunately, studies of related proteins from other species containing both BEACH and WD40 domains do not shed much light on the function of Lyst. The functions of such proteins appear to be quite diverse ( 4). Studies of the LvsB protein of D. Discoideum, the closest BEACH- and WD40-containing homolog of Lyst in this species, indicate that LvsB in fact negatively regulates lysosome fusion ( 35;36). LvsB-null mutants contain enlarged vesicles that appear to be acidic lysosomes, and display increased fusion rates ( 35). A common mechanism or function shared by all BEACH- and WD40-containing proteins remains to be discovered.
Reports suggest that Lyst may regulate protein kinase C (PKC) activity. Membrane-bound PKC activity is reportedly abnormally downregulated in beige fibroblasts, macrophages and polymorphonuclear leukocytes after phorbol ester treatment ( 37). This aberrant PKC downregulation, and the formation of giant granules, can be inhibited by treatment with the thiol proteinase inhibitor E-64-d, a calpain inhibitor ( 38). The authors suggest that inhibition of calpain-mediated PKC proteolysis prevents giant granule formation ( 38). The mechanism of PKC regulation of lysosomes is unknown.
The souris mutation may result in expression of a significantly truncated protein missing four of the ARM/HEAT motifs, as well as the BEACH and WD40 domains. Although the exact function of these domains remain to be discovered, it is likely that the souris mutation results in a protein with severely affected function.
|
|---|
| Primers |
Primers cannot be located by automatic search.
|
|---|
| Genotyping | Souris genotyping is performed by amplifying the region containing the mutation using PCR, followed by sequencing of the amplified region to detect the single nucleotide change.
Primers for PCR amplification
Souris(F): 5’- TCACCCTTCATGTAGACCAGGAACC -3’
Souris(R): 5’- TGCAGGGCCAAAGCCATATCTAAAC -3’
PCR program
1) 94°C 2:00
2) 94°C 0:30
3) 56°C 0:30
4) 72°C 1:00
5) repeat steps (2-4) 29X
6) 72°C 7:00
7) 4°C ∞
Primers for sequencing
Souris_seq(F): 5’- CAGAAGTCAATGTGCCTTACTTGC -3’
Souris_seq(R): 5’- GATGACCTTGAAATTCTGACCATCC -3’
The following sequence of 1228 nucleotides (from Genbank genomic region NC_000079 for linear genomic sequence of Lyst) is amplified:
92341 tcacccttca tgtagaccag gaacctgtgt gagtgttctc tgtgtatctt gtgttaatct
92401 ctagctgttg tttatgttgc tctcttaacc tgtatctttc tctacaagac tgtattttca
92461 acagaagtca atgtgcctta cttgcattcc tagtgggcag ggtatattgc ttgctgtata
92521 gagtaaacat gaacagttct taaatcattg catgctggta gcgtagagtt taaccagttc
92581 tttgtctttt aatttttcgg ctgttttgct tatagtatct tctgttttgt tttggactca
92641 ctttagcatt cctgtgaacg aatacaaatt gctcgcatgt gatatacagc agcttttcat
92701 agcagttaca attcatgctt gcagttcctc aggcacacag tattttagag tgattgaaga
92761 ccttattgta cttcttggat atcttcataa tagcaaaaac aagaggacac aaagtaagaa
92821 tggattttaa tggaattttt ctcataactt ttactgaaga tttaaagtta attatgtgtt
92881 ggtattaatt ggtatttaaa gcaaccttta aattgttgaa aatagagtga ttgatggttt
92941 gatgtaatta gtagttggca atacatcact ggcagaaact agtaattttt tttaccaagt
93001 agaatacaaa ggcatatgtt ggaagaataa tgggaatgct agcgtgactc atgagtgaaa
93061 gcgtgtaggg tattggagct tttgctttat acaaagttat tggtttttga aacagtcatt
93121 taaagtgaaa tcagtttagg atttatcatt ttttctattc aaatttttat agttgtagct
93181 agatatggtg atacatattc aagagatgga ggcaggatgg tcagaatttc aaggtcatct
93241 tcagctacat agcaggctaa cctgggctac aggaaacatg tctcaaaaat atcaaaaaat
93301 ttgtagtgac catatgggaa tttattttaa cattattttt aaatgtagct tagaaacact
93361 aacatcaaga acccatgttg atgaaacaat tttaacttgt gtctaaggcc ttttcattta
93421 gatattgaat attatagctt gaatattagg ttatatgcct tagatgttag ataatatatt
93481 aaataagttt agttaaaaga gttaaacaat tatgatagct tgaaaacatt ctagtctctt
93541 tgggtttaga tatggctttg gccctgca
Primer binding sites are underlined; sequencing primer binding sites are highlighted in gray; the mutated T is shown in red.
|
|---|
| References | 1. Barbosa, M. D., Nguyen, Q. A., Tchernev, V. T., Ashley, J. A., Detter, J. C., Blaydes, S. M., Brandt, S. J., Chotai, D., Hodgman, C., Solari, R. C., Lovett, M., and Kingsmore, S. F. (1996) Identification of the homologous beige and Chediak-Higashi syndrome genes, Nature 382, 262-265. 2. Perou, C. M., Moore, K. J., Nagle, D. L., Misumi, D. J., Woolf, E. A., McGrail, S. H., Holmgren, L., Brody, T. H., Dussault, B. J., Jr., Monroe, C. A., Duyk, G. M., Pryor, R. J., Li, L., Justice, M. J., and Kaplan, J. (1996) Identification of the murine beige gene by YAC complementation and positional cloning, Nat. Genet. 13, 303-308. 3. Nagle, D. L., Karim, M. A., Woolf, E. A., Holmgren, L., Bork, P., Misumi, D. J., McGrail, S. H., Dussault, B. J., Jr., Perou, C. M., Boissy, R. E., Duyk, G. M., Spritz, R. A., and Moore, K. J. (1996) Identification and mutation analysis of the complete gene for Chediak-Higashi syndrome, Nat. Genet. 14, 307-311. 5. Andrade, M. A., Petosa, C., O'Donoghue, S. I., Muller, C. W., and Bork, P. (2001) Comparison of ARM and HEAT protein repeats, J Mol. Biol. 309, 1-18. 8. Miura, S., Gan, J. W., Brzostowski, J., Parisi, M. J., Schultz, C. J., Londos, C., Oliver, B., and Kimmel, A. R. (2002) Functional conservation for lipid storage droplet association among Perilipin, ADRP, and TIP47 (PAT)-related proteins in mammals, Drosophila, and Dictyostelium, J Biol. Chem. 277, 32253-32257. 9. Ward, D. M., Shiflett, S. L., Huynh, D., Vaughn, M. B., Prestwich, G., and Kaplan, J. (2003) Use of expression constructs to dissect the functional domains of the CHS/beige protein: identification of multiple phenotypes, Traffic. 4, 403-415. 10. Sondek, J., Bohm, A., Lambright, D. G., Hamm, H. E., and Sigler, P. B. (1996) Crystal structure of a G-protein beta gamma dimer at 2.1A resolution, Nature 379, 369-374. 11. Barbosa, M. D., Barrat, F. J., Tchernev, V. T., Nguyen, Q. A., Mishra, V. S., Colman, S. D., Pastural, E., Dufourcq-Lagelouse, R., Fischer, A., Holcombe, R. F., Wallace, M. R., Brandt, S. J., de Saint, B. G., and Kingsmore, S. F. (1997) Identification of mutations in two major mRNA isoforms of the Chediak-Higashi syndrome gene in human and mouse, Hum. Mol. Genet. 6, 1091-1098. 12. Perou, C. M., Leslie, J. D., Green, W., Li, L., Ward, D. M., and Kaplan, J. (1997) The Beige/Chediak-Higashi syndrome gene encodes a widely expressed cytosolic protein, J Biol. Chem. 272, 29790-29794. 13. Faigle, W., Raposo, G., Tenza, D., Pinet, V., Vogt, A. B., Kropshofer, H., Fischer, A., de Saint-Basile, G., and Amigorena, S. (1998) Deficient peptide loading and MHC class II endosomal sorting in a human genetic immunodeficiency disease: the Chediak-Higashi syndrome, J Cell Biol. 141, 1121-1134. 15. Zhao, H., Boissy, Y. L., bdel-Malek, Z., King, R. A., Nordlund, J. J., and Boissy, R. E. (1994) On the analysis of the pathophysiology of Chediak-Higashi syndrome. Defects expressed by cultured melanocytes, Lab Invest 71, 25-34. 20. Abo, T., Roder, J. C., Abo, W., Cooper, M. D., and Balch, C. M. (1982) Natural killer (HNK-1+) cells in Chediak-Higashi patients are present in normal numbers but are abnormal in function and morphology, J Clin. Invest 70, 193-197. 21. Roder, J. C., Haliotis, T., Klein, M., Korec, S., Jett, J. R., Ortaldo, J., Heberman, R. B., Katz, P., and Fauci, A. S. (1980) A new immunodeficiency disorder in humans involving NK cells, Nature 284, 553-555. 22. Rubin, C. M., Burke, B. A., McKenna, R. W., McClain, K. L., White, J. G., Nesbit, M. E., Jr., and Filipovich, A. H. (1985) The accelerated phase of Chediak-Higashi syndrome. An expression of the virus-associated hemophagocytic syndrome? Cancer 56, 524-530. 23. Haddad, E., Le, D. F., Blanche, S., Benkerrou, M., Rohrlich, P., Vilmer, E., Griscelli, C., and Fischer, A. (1995) Treatment of Chediak-Higashi syndrome by allogenic bone marrow transplantation: report of 10 cases, Blood 85, 3328-3333. 24. Kazmierowski, J. A., Elin, R. J., Reynolds, H. Y., Durbin, W. A., and Wolff, S. M. (1976) Chediak-Higashi syndrome: reversal of increased susceptibility to infection by bone marrow transplantation, Blood 47, 555-559. 25. Sung, J. H., Meyers, J. P., Stadlan, E. M., Cowen, D., and Wolf, A. (1969) Neuropathological changes in Chediak-Higashi disease, J Neuropathol. Exp. Neurol. 28, 86-118. 27. Kelly, E. M. (1957) Beige, bg, Mouse News Lett 16, 36.
33. Tchernev, V. T., Mansfield, T. A., Giot, L., Kumar, A. M., Nandabalan, K., Li, Y., Mishra, V. S., Detter, J. C., Rothberg, J. M., Wallace, M. R., Southwick, F. S., and Kingsmore, S. F. (2002) The Chediak-Higashi protein interacts with SNARE complex and signal transduction proteins, Mol. Med. 8, 56-64. 34. Huynh, C., Roth, D., Ward, D. M., Kaplan, J., and Andrews, N. W. (2004) Defective lysosomal exocytosis and plasma membrane repair in Chediak-Higashi/beige cells, Proc. Natl. Acad. Sci. U. S. A 101, 16795-16800. 35. Harris, E., Wang, N., Wu Wl, W. L., Weatherford, A., De, L. A., and Cardelli, J. (2002) Dictyostelium LvsB mutants model the lysosomal defects associated with Chediak-Higashi syndrome, Mol. Biol. Cell 13, 656-669. 37. Ito, M., Tanabe, F., Takami, Y., Sato, A., and Shigeta, S. (1988) Rapid down-regulation of protein kinase C in (Chediak-Higashi syndrome) beige mouse by phorbol ester, Biochem. Biophys. Res. Commun. 153, 648-656. |
|---|
| Science Writers | Eva Marie Y. Moresco |
|---|
| Illustrators | Diantha La Vine, Katherine Timer |
|---|
| Authors | Sophie Rutschmann, Celine Eidenschenk, Bruce Beutler |
|---|


 The souris mutation was mapped to Chromosome 13, and corresponds to a T to A transversion in the donor splice site of intron 27 (GTAAG -> GAAAG) of the Lyst gene (position 92815 in the Genbank genomic region NC_000079 for linear genomic DNA sequence of Lyst). As Lyst cDNA from souris mice has not been sequenced, it is unknown how the mutation affects processing of the Lyst transcript. The mutation likely results in skipping of the 167-nucleotide exon 27 (out of 53 total exons), destroying the reading frame (aberrant amino acids after position 2474), and creating a premature stop codon that would truncate the protein after amino acid 2482:
The souris mutation was mapped to Chromosome 13, and corresponds to a T to A transversion in the donor splice site of intron 27 (GTAAG -> GAAAG) of the Lyst gene (position 92815 in the Genbank genomic region NC_000079 for linear genomic DNA sequence of Lyst). As Lyst cDNA from souris mice has not been sequenced, it is unknown how the mutation affects processing of the Lyst transcript. The mutation likely results in skipping of the 167-nucleotide exon 27 (out of 53 total exons), destroying the reading frame (aberrant amino acids after position 2474), and creating a premature stop codon that would truncate the protein after amino acid 2482:
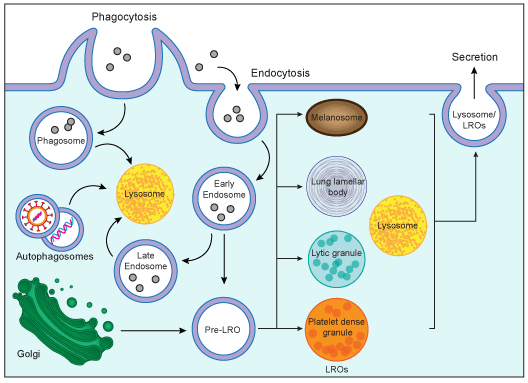 Figure 2. Biogenesis of lysosomes and lysosome-relateded organelles (LORs). Lysosomes are cytoplasmic organelles responsible for degradation within a cell. LROs share some features with lysosomes but have distinct morphologies and functions.
Figure 2. Biogenesis of lysosomes and lysosome-relateded organelles (LORs). Lysosomes are cytoplasmic organelles responsible for degradation within a cell. LROs share some features with lysosomes but have distinct morphologies and functions.